Integration of Ex-vivo precision-cut liver slice (PCLS) culture with microfluidic NMR metabolomics
Seminar Abstract
Bishnubrata Patra(1), Manvendra Sharma(1), Ruby Karsten (2), Maciej Grajewski (2), Sabeth Verpoorte (2), and Marcel Utz (1)
1 School of Chemistry, University of Southampton, UK
2 School of Pharmacy, University of Groningen, The Netherlands
ABSTRACT
We present a novel microfluidic perfusion system for liver tissue slices that allows direct characterization of the perfusion fluid by micro nuclear magnetic resonance (NMR) spectroscopy and imaging. This system enables direct non-invasive observation of the metabolic processes on the liver slice in real time. Integration of a microfluidic system with high-performance NMR spectroscopy has been achieved with careful management of sensitivity and magnetic susceptibility effects to maintain spectral resolution. The system presented here combines excellent NMR
performance with the ability to sustain the PCLS over several hours of perfusion.
KEYWORDS: Tissue slice culture, Microfluidics, Metabolomics, NMR
INTRODUCTION
Ex-vivo culture of tissue provides an alternative to animal models to study the effect of external factors like exposure to drugs, environmental changes, infection, and inflammation, requiring significantly less resources [1] and providing more detailed information. Observation of metabolism
by NMR is well established. We present a microfluidic perfusion culture system for PCLS that allows direct NMR observation of the metabolite composition of the perfusate after contact with the slice using a home made transmission-line NMR probe [2].
EXPERIMENTAL
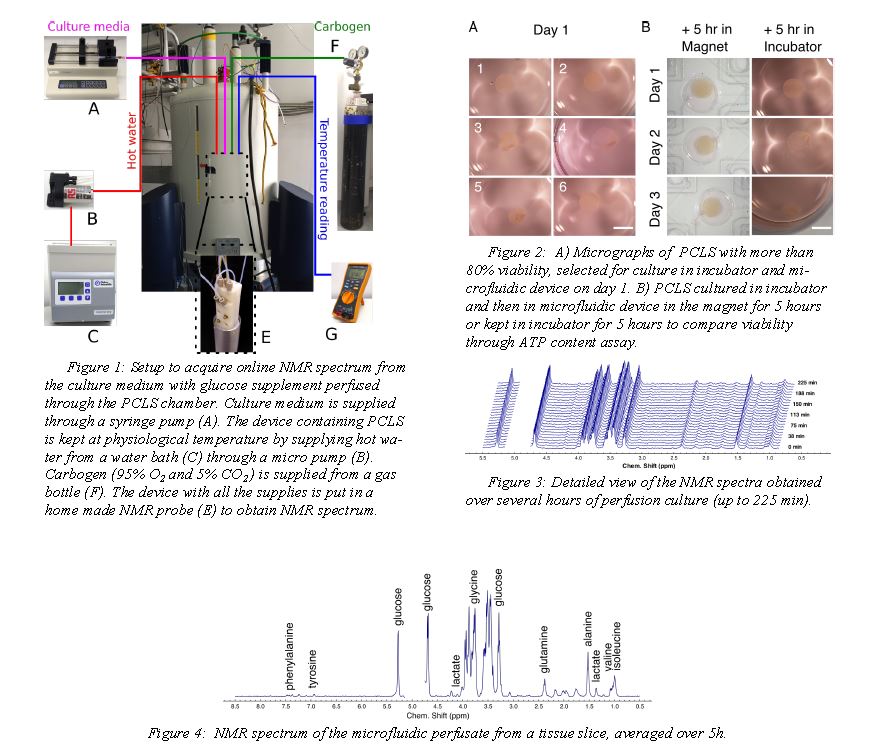
The NMR setup and the perfusion chip are shown in figure 1. The microfluidic device (figure 1E) with the PCLS is sandwiched between polydimethylsiloxane(PDMS) membranes and chip holders. Culture
medium is supplied with a syringe pump at a flow rate of 8 µl/min. Both oxygen and carbon dioxide are exchanged by diffusion through the PDMS membrane. The device containing PCLS is kept at physiological temperature bysupplying hot water from a water bath (figure 1C) through a micro pump ( figure 1B).
RESULTS AND DISCUSSION
Murine PCLS were incubated in well plates at 37°C in 80% O2 and 5% CO2 after slicing. Their viability was ascertained by LDH leakage assay before the perfusion experiment at day 1. After 1-3 days of incubation, they were transferred into the perfusion system. NMR spectra were continuously
recorded for up to 5 hours of perfusion. Incremental NMR spectra obtained over the course of the perfusion demonstrates stability of the system over several hours (figure 3). The proton NMR spectrum in Figure 4, averaged over the duration of the experiment, gives an indication of the
quality of the spectral information. At least 20 different metabolites are clearly apparent from this spectrum; most prominently, glucose, alanine, glutamine, glutamate, valine, leucine, and isoleucine. A small lactate peak is visible t 1.32~ppm. After 5h of perfusion, viability of the slices was
verified using ATP content/ µg of protein assay.
CONCLUSION
These results demonstrate that PCLS can be kept viable in an NMR-compatible microfluidic system, while high-quality NMR data of the perfusate is extracted. This opens the way for further studies to explore the
metabolic activity of slices exposed to various external stimuli.
ACKNOWLEDGEMENTS
This research project is funded by the horizon 2020 Framework program of the European Union. The authors sincerely acknowledge the discussion with Prof. Peter Olinga regarding tissue slice culture.
REFERENCES
[1] I. A. M. de Graaf, P. Olinga, M. H. de Jager, M. T. Merema, R. d Kanter, E. G. Van de Kerkhof, G. M. M Groothuis, Preparation and Incubation of Precision-Cut Liver and Intestinal Slices for Application in Drug
Metabolism and Toxicity Studies. Nature Protocols 2010, 5 (9), 1540-1551.
[2] M. Sharma, M. Utz, Modular Transmission Line Probes for Microfluidic Nuclear Magnetic Resonance Spectroscopy and Imaging. Journal of Magnetic Resonance 2019, 303, 75-81.
